|
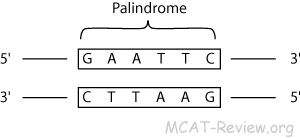
10/31/2022 0 Comments Palindromic sequence Take a look at the example,Ī cruciform hairpin DNA, formed by a palindromic sequence. Near-palindromic sequences which are not exactly similar, as aforementioned, like the inverted repeats, commonly have cruciform hairpins due to having larger spacer elements within the palindromic elements. Studies evidence that cruciform hairpins are regions more prone to DNA breakage, deletion and translocations which are inherited as well. They form a hairpin-like secondary structure by folding back and forming a structure known as “cruciform”. Henceforth, longer palindrome or near-palindrome sequences are not so suitable for our genome as they can form a hairpin. In addition, their paper also suggests that palindromes lower than 50bp prevent genomic DNA degradation, on the other side, longer sequences than 50bp create genome instability thereby a disease. Ganapathiraju et al., 2020 Explained in their paper entitled “A reference catalog of DNA palindromes in the human genome and their variations in 1000 genomes” that 30% of such sequences are prone to variations and are involved in single and multi-trait disorders. Studies suggest that the length of a palindromic sequence has a serious role in various genetic activities and is also involved in deletions, insertions or diseases.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
